SatTCR pipeline
SatTCR is a Snakemake pipeline for assembling T Cell Receptor (TCR) repertoire data by using MIXCR, and perform a saturation analysis.
TCR profiling
Profiling the TCR repertoire using next-generation sequencing (NGS) to quantify adaptive immune responses has become common in human and animal research. TCR counts from NGS data provide a way to quantify T cell response to vaccines, cancer, or infectious diseases for preclinical and clinical health studies.
SatTCR modules
SatTCR is composed by 4 modules described in the image below:
Quality control: QC diagnostics of the sequencing data.
Clonotype assembly: Assembly of the TCR clonotypes using MIXCR
Saturation analysis:: Sequential bootstrapping of the sequencing data to assembly clonotypes with a fixed # of sequencing reads.
Report of SatTCR results: The report contains data visualization of statistics summarizing the TCR repertoire assembled with the complete data and sampled version of the sequencing files.
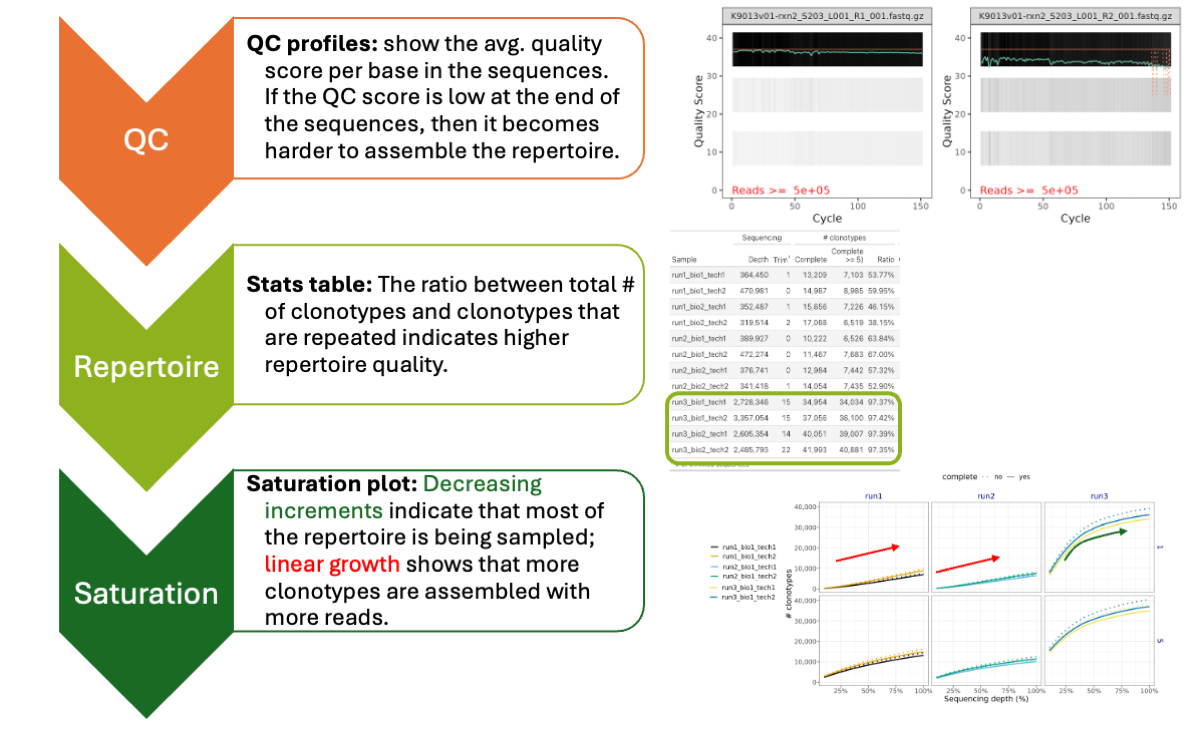
How interpret SatTCR results?
A SatTCR provides multiple analysis. The key points to pay attention are: